Coccinella
What is coccinella?
Coccinella is an innovative open-source framework developed for high-throughput behavioral analysis. Leveraging the power of distributed microcomputers, it facilitates real-time tracking of small animals, such as Drosophila melanogaster. Complementing this tracking capability, coccinella employs advanced statistical learning techniques to decipher and categorize observed behaviors. Unlike many high-resolution systems that often require significant resources and may compromise on throughput, coccinella strikes a balance, offering both precision and efficiency. Built upon the foundation of ethoscopes, this platform extracts minimalist yet crucial information from behavioral paradigms. Notably, in comparative studies, coccinella has demonstrated superior performance in recognizing pharmacobehavioral patterns, achieving this at a fraction of the cost of other state-of-the-art systems. This framework promises to complement current ethomics tools by providing a cost-effective, efficient, and precise tool for behavioral research. Coccinella analysis can be done in ethoscopy, a Python framework for analysis of ethoscope data.
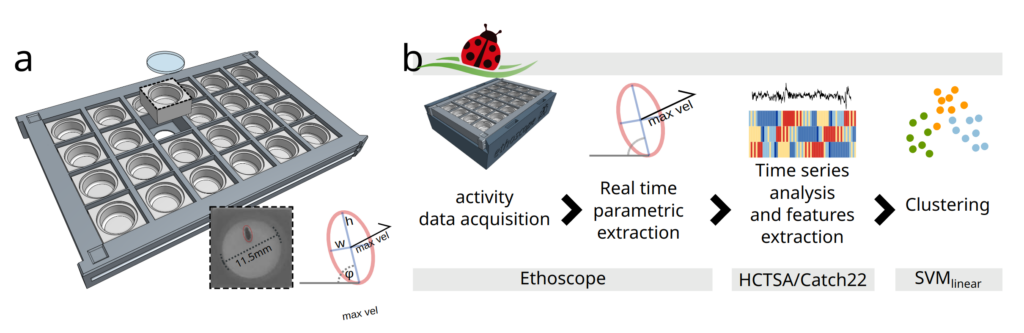
How does it work?
Coccinella uses ethoscopes to extract information about the activity of flies in real time. Ethoscopes are machines that use distributed computing via Raspberry PI to detect and interfere with behaviour. Given the off-the-shelf nature of the devices, the all setup is inexpensive and scales up very easily. As term of reference: our lab currently employs about 100 ethoscopes, with a processing power of 20 flies each.
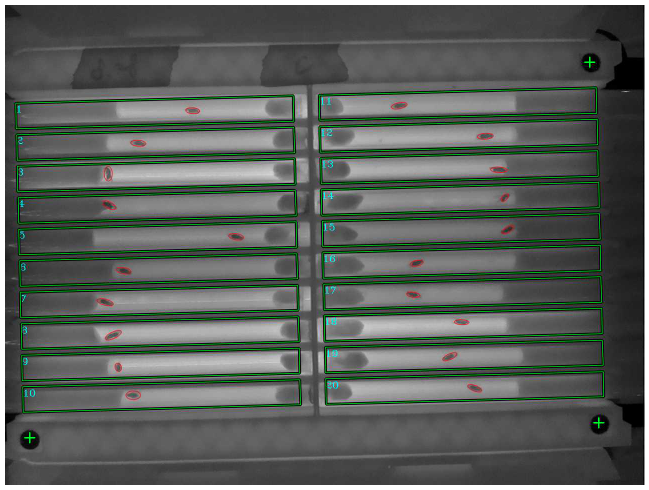

Data about the activity of the animal are then fed to a high-throughput toolbox for time series analysis called HCTSA or Catch22, initially developed at Imperial College London by our colleagues in the Maths Department. The toolbox performs numerous statistical tests aimed at segregating data in an unsupervised way and can therefore be used to cluster together data that the machine recognises as similar. In our case, we tried to identify which drugs have similar mode of action, potentially recognising and assigning the appropriate pharmacological pathways to new, uncharacterized compounds.
We also compared the performance of coccinella to state of the art systems and found that it performs even better!
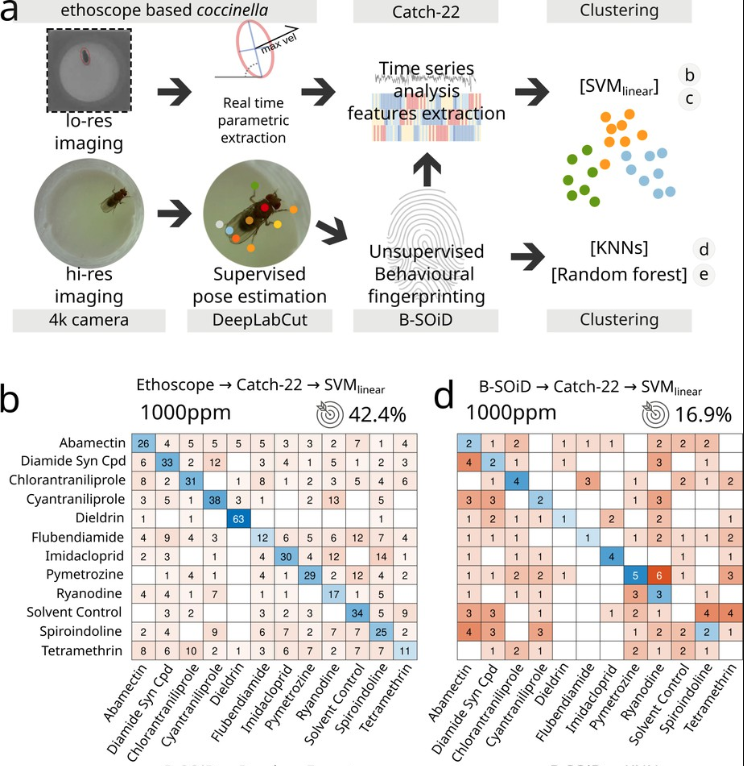
This is important from the technical point of view but also from the standpoint of neuroscience because it shows that “less is more” when it comes to extracting and recognising behavioural data. In other words, you don’t need to carefully label posture and movement when characterising behaviour: reducing activity to its minimal terms actually works even better!
 | Coccinella paper on eLife |
 | Ethoscopy / Ethoscope-lab paper on Bioinformatics Advances |
 | Ethoscopy on GitHub |
 | Ethoscopy on PyPi |
| Ethoscope-lab Docker container on DockerHub | |
 | Jupyter Notebook tutorials for Ethoscopy on GitHub |
 | Ethoscopy and Ethoscope-lab documentation on bookstack |
| Raw data and all notebooks reproducing the paper’s figures on Zenodo |

Comments are Disabled